|
FiveFold: Conformation Ensemble of Structures for Protein Structure Prediction |
a FiveFold approach was developed with a single sequence method. It applied the protein folding shape code (PFSC) uniformly to expose local folds of five amino acid residues, formed the protein folding variation matrix (PFVM) to reveal local folding variations along sequence, obtained a massive number of folding conformations in PFSC strings, and then an ensemble of multiple conformational protein structures is constructed. The red arrows indicated the process to predict an ensemble of protein structures from a protein sequence with the PFVM.
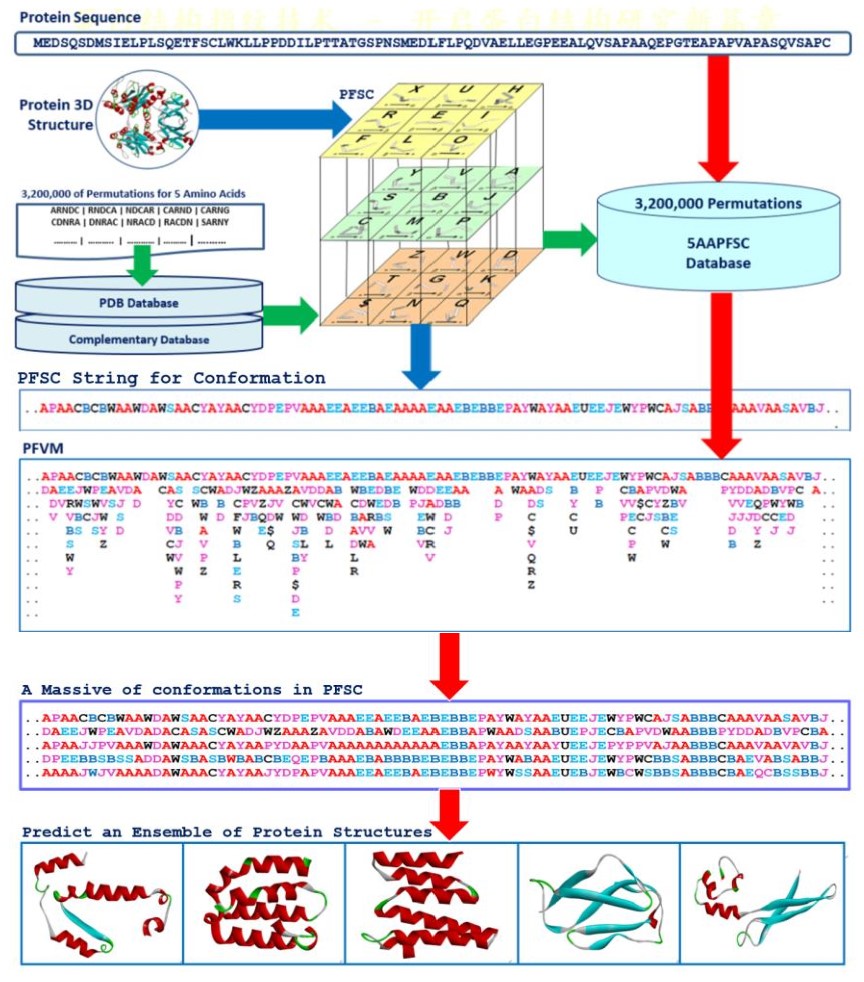
Structure Prediction for DNA-binding domain of P53_HUMAN
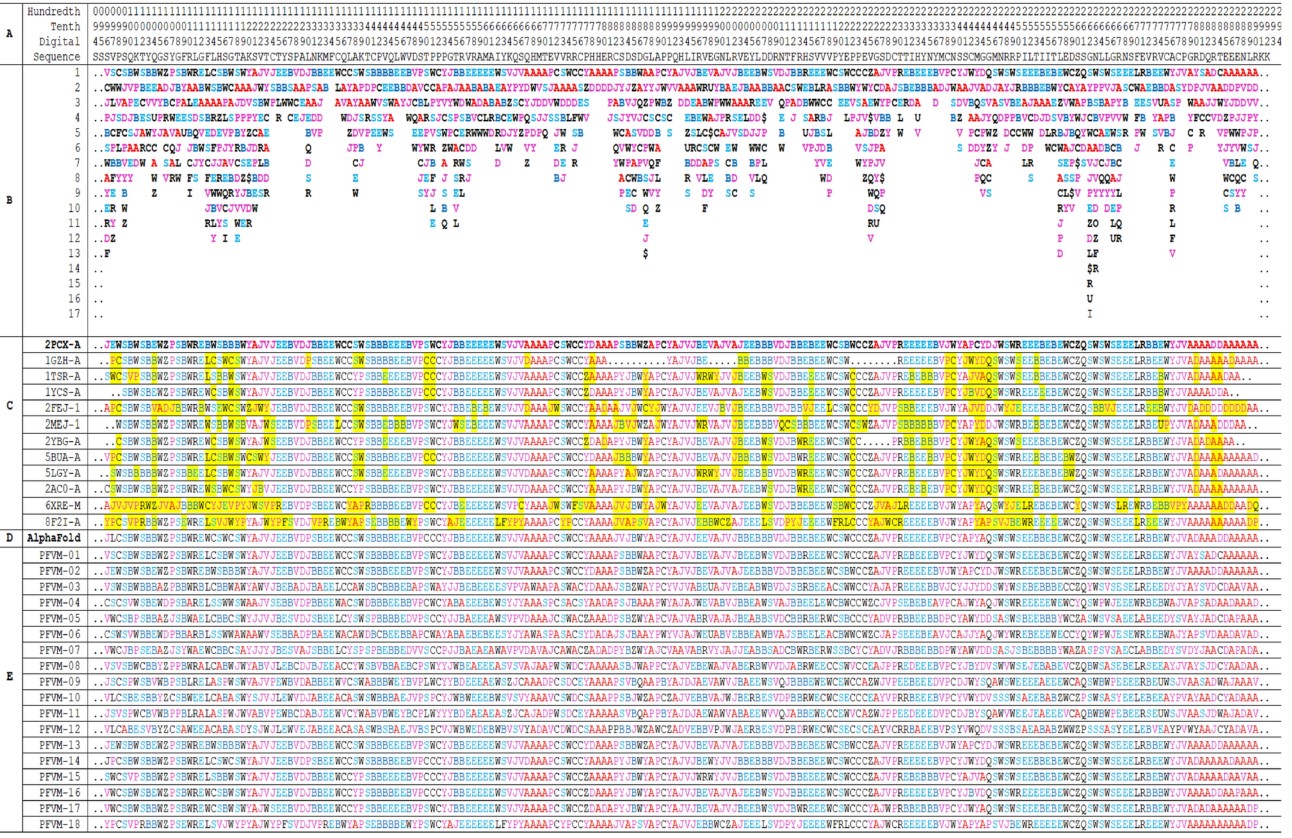 The PFVM and PFSC for DNA-binding domain region (100–288) of P53_HUMAN (P04637).
The protein sequence is listed in section A; the PFVM in section B; the conformations in PFSC string for
given 3D structures in section C; the conformation of AlphaFold in PFSC string in section D and a set of
conformations in PFSC string predicted by FiveFold in section E. The PFSC letters with red and pink colors are
for typical helix and alike-helix local folds; the blue and light blue colors for beta strand and alike-beta strand
and block color for irregular folds.
The PFVM and PFSC for DNA-binding domain region (100–288) of P53_HUMAN (P04637).
The protein sequence is listed in section A; the PFVM in section B; the conformations in PFSC string for
given 3D structures in section C; the conformation of AlphaFold in PFSC string in section D and a set of
conformations in PFSC string predicted by FiveFold in section E. The PFSC letters with red and pink colors are
for typical helix and alike-helix local folds; the blue and light blue colors for beta strand and alike-beta strand
and block color for irregular folds.
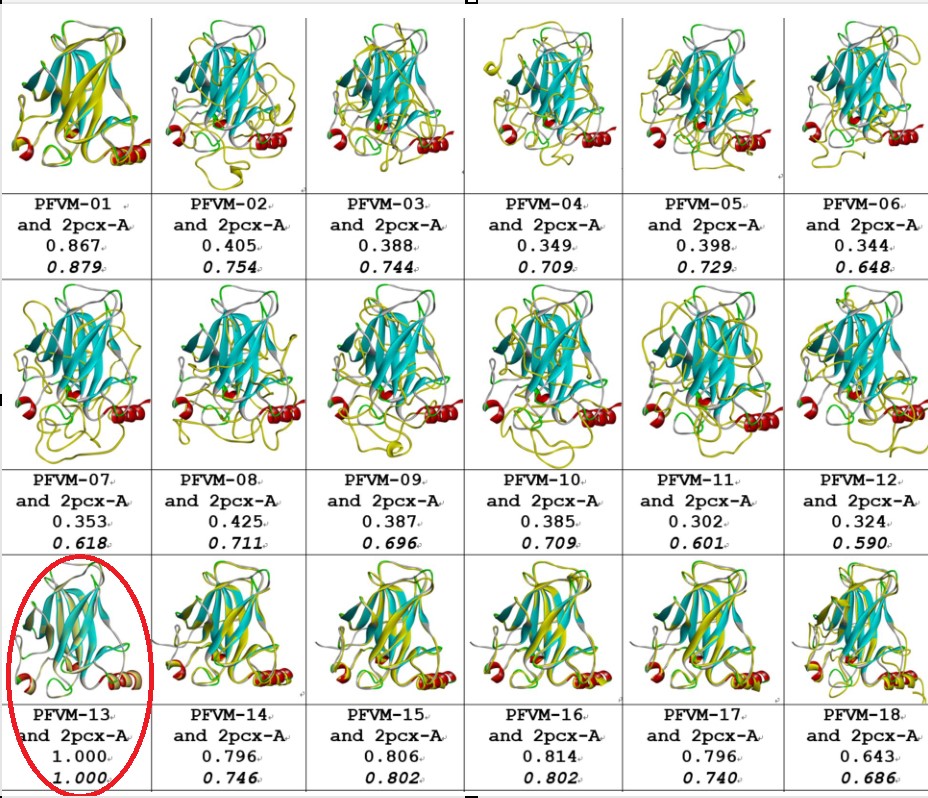
Comparisons between the given 2PCX-A structure and 18 of FiveFold predicted structure (yellow) for DNA-binding domain (100–288) of P53_HUMAN. The images of superposition of each pair of structures are displayed. The conformation name for FiveFold and given structure are listed under image. The overlap similarity score is displayed under structure name, which was obtained by overlaying the molecule structures with alignment by a combination of 50% steric component and 50% electrostatic component in software of Discovery Studio v24.1.0. Also, with PFSC string alignment, the score of protein folding structural alignment (PFSA) in italics is listed.
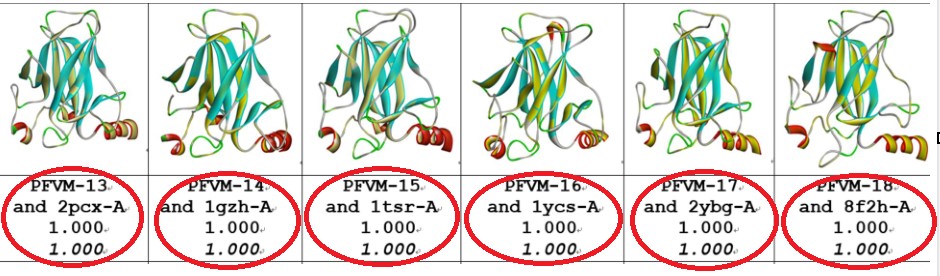 Comparisons between the FiveFold predicted structure (yellow) and each of 6 given structures for
DNA-binding domain (100–288) of P53_HUMAN. The images of superposition of each pair of structures
are displayed. The conformation name for FiveFold and given structure are listed under image. The overlap
similarity score is displayed under structure name, which was obtained by overlaying the molecule structures
with alignment by a combination of 50% steric component and 50% electrostatic component in software of
Discovery Studio v24.1.0. Also, with PFSC string alignment, the score of protein folding structural alignment
(PFSA) in italics is listed.
Comparisons between the FiveFold predicted structure (yellow) and each of 6 given structures for
DNA-binding domain (100–288) of P53_HUMAN. The images of superposition of each pair of structures
are displayed. The conformation name for FiveFold and given structure are listed under image. The overlap
similarity score is displayed under structure name, which was obtained by overlaying the molecule structures
with alignment by a combination of 50% steric component and 50% electrostatic component in software of
Discovery Studio v24.1.0. Also, with PFSC string alignment, the score of protein folding structural alignment
(PFSA) in italics is listed.
References:
